|
 |
- Search
| J Stroke > Volume 17(1); 2015 > Article |
Abstract
Cerebral small vessel disease (SVD) is an important cause of stroke and cognitive impairment among the elderly and is a more frequent cause of stroke in Asia than in the US or Europe. Although traditional risk factors such as hypertension or diabetes mellitus are important in the development of cerebral SVD, the exact pathogenesis is still uncertain. Both, twin and family history studies suggest heritability of sporadic cerebral SVD, while the candidate gene study and the genome-wide association study (GWAS) are mainly used in genetic research. Robust associations between the candidate genes and occurrence of various features of sporadic cerebral SVD, such as lacunar infarction, intracerebral hemorrhage, or white matter hyperintensities, have not yet been elucidated. GWAS, a relatively new technique, overcomes several shortcomings of previous genetic techniques, enabling the detection of several important genetic loci associated with cerebral SVD. In addition to the more common, sporadic cerebral SVD, several single-gene disorders causing cerebral SVD have been identified. The number of reported cases is increasing as the clinical features become clear and diagnostic examinations are more readily available. These include cerebral autosomal dominant arteriopathy with subcortical infarcts and leukoencephalopathy, cerebral autosomal recessive arteriopathy with subcortical infarcts and leukoencephalopathy, COL4A1-related cerebral SVD, autosomal dominant retinal vasculopathy with cerebral leukodystrophy, and Fabry disease. These rare single-gene disorders are expected to play a crucial role in our understanding of cerebral SVD pathogenesis by providing animal models for the identification of cellular, molecular, and biochemical changes underlying cerebral small vessel damage.
Cerebral small vessel disease (SVD) is an important cause of stroke, cognitive impairment, and mood disorder among the elderly.1 It accounts for 15% to 26% of ischemic stroke in the US and Europe.2,3,4,5,6 In Asia, cerebral SVD is responsible for an even greater proportion of ischemic stroke, ranging from 25% to 54%.7,8,9,10,11 In addition, it is the most important cause of intracerebral hemorrhage (ICH)12 and contributes to a substantial proportion of vascular dementia.13
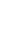
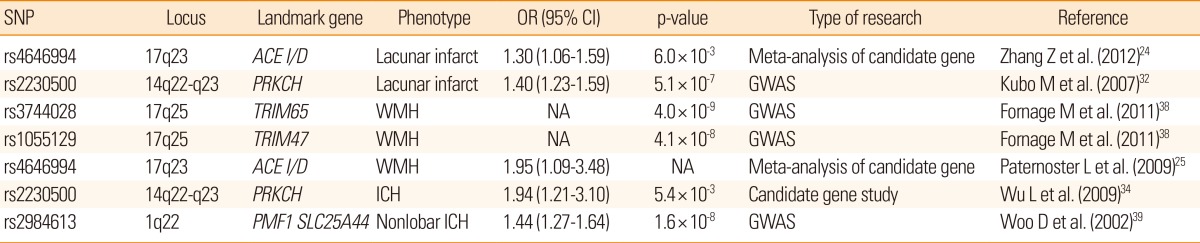
Although traditional risk factors are important in the development of cerebral SVD14 the exact cause is still uncertain because not all individuals with those risk factors develop the disease and often, known risk factors are absent in patients with cerebral SVD.15 Ischemic stroke is possibly a complex genetic disorder in which interactions between candidate genes and environmental factors or established risk factors are important in the development of clinical phenotypes.16 Therefore, genetics may play an important role in understanding the pathophysiology of sporadic cerebral SVD. With advances in genetic methodology such as genome-wide association study (GWAS) and meta-analysis of individual candidate gene study, both, extensive and in-depth information on the susceptible genetic loci associated with sporadic cerebral SVD (Table 1) is now available. In addition, several single-gene disorders causing cerebral SVD have been discovered, including cerebral autosomal dominant arteriopathy with subcortical infarcts and leukoencephalopathy (CADASIL), cerebral autosomal recessive arteriopathy with subcortical infarcts and leukoencephalopathy (CARASIL), COL4A1-related cerebral SVD, autosomal dominant retinal vasculopathy with cerebral leukodystrophy (RVCL), and Fabry disease (FD; Table 2). Recognition of the genetic aspect of cerebral SVD could lead to improved diagnosis and treatment of these rare single-gene disorders as well as sporadic cerebral SVD. In this review, we have focused on the recent advances in genetics of both, sporadic and familial forms of cerebral SVD.
Twin studies have suggested that monozygotic twins were more likely to be concordant for risk of stroke than dizygotic twins and a positive family history was a risk factor for stroke in both case-control and cohort studies.17 For subtypes of ischemic stroke, a family history of stroke was a significant risk factor for large vessel disease (odds ratio [OR], 2.24 to 2.54) and for SVD (OR, 1.93 to 2.62) but not for cardioembolic stroke.18,19 Cerebral white matter hyperintensities (WMH) are one of the main imaging features of SVD, and they are frequently found in patients with lacunar stroke on brain magnetic resonance imaging (MRI).14 Twin studies also indicate a genetic contribution to WMH with concordance rates of 61% in monozygotic pairs and that of 38% in dizygotic pairs for large proportion of WMH.20 The findings were also replicated in the family-based Framingham study, with an overall heritability of approximately 52% to 78%.21 ICH, especially nonlobar ICH, results from the rupture of small penetrating arteries of the brain and is another important clinical manifestation of cerebral SVD.12 Heritability of nonlobar or primary ICH was suggested by case-control studies showing associations between family history and an increased risk of ICH (adjusted OR, 2.1 to 6.3).18,22
Candidate gene study is an approach involving exploring the genetic influences on a complex trait by identifying candidate genes that might have a role in the etiology of the disease and this method has been widely used for the study of complex diseases.23 In a comprehensive meta-analysis of 187 candidate genetic polymorphism in 37,481 stroke patients, 6 polymorphisms, namely, factor V Leiden, angiotensin converting enzyme (ACE) insertion/deletion (I/D), methylene tetrahydrofolate reductase (MTHFR) C677T, prothrombin G20210A, plasminogen activator inhibitor-1 4G/5G allele, and glycoprotein IIIa L33P, were found to have an association with increased risk of stroke. Among these, ACE I/D polymorphism was more specifically associated with increased risk of lacunar stroke than with other types of ischemic stroke. In another meta-analysis of 50 case-control studies, homozygote DD of ACE gene polymorphism showed a 37% higher risk of ischemic stroke compared with homozygote II and heterozygote ID, and a subgroup analysis showed that the risk was more prominent among Asians and SVD subtype of ischemic stroke.24 Researchers have also looked for associations between polymorphisms in various candidate genes and WMH in brain images. In a meta-analysis including 19 candidate gene polymorphisms from 19,000 subjects, no evidence of association was found between apolipoprotein E (APOE) (ε4+/-), MTHFR C677T, or angiotensinogen M235T, and WMH.25 Only ACE I/D polymorphism showed a significant association (OR, 1.95; 95% CI, 1.09 to 3.48) with the WMH.
Candidate genes that have been investigated most frequently for ICH are APOE, plasminogen activator inhibitor-1 4G/5G, ACE/ID, factor V Leiden, MTHFR C677T, and factor XIII V34L polymorphism.26 In a meta-analyses that included 6,359 cases derived from 55 case-control studies, homozygous for the ACE/I allele and for the 5G allele in plasminogen activator inhibitor-1 4G/5G polymorphism were found to have statistically significant associations with hemorrhagic stroke (including ICH, subarachnoid hemorrhage, vascular malformation, and cerebral aneurysm) when they were analyzed together, but not specifically for ICH. A large-scale genetic association study with 2,189 ICH cases and 4041 controls found that both, the ε2 and the ε4 alleles of APOE were significantly associated with increased risk of lobar ICH. Although, an association was also found between nonlobar ICH and the ε4 allele, the association did not reach predefined significance.27 In a subsequent research, the APOE ε2 allele was also associated with larger ICH volumes, increased mortality, and poorer functional outcomes than non-carriers for patients with lobar ICH.28 In a meta-analyses including 10 studies with 7,351 patients, the APOE ε4 allele was associated with increased risk of cerebral microbleeds in any location compared with common ε3/ε3 genotype (pooled OR 1.22, 95% confidence interval [CI] 1.05 to 1.41, P=0.01), and the association was stronger for strictly lobar, cerebral microbleeds.29 The biological role of APOE genotype in the development of nonlobar ICH or cerebral microbleeds needs further exploration because cerebral amyloid angiopathy-related vasculopathy rarely occurs in vessels of deep gray matter.30
In summary, although meta-analyses of candidate gene studies have overcome an important limitation of small sample size in case of individual case-control studies, these findings should be interpreted with caution, as the results may be confounded by publication bias.
GWAS is a new method to study common genetic variation across the entire human genome, designed to identify genetic associations with observable traits.31 It is particularly useful to investigate common diseases with complex traits such coronary artery disease and stroke. Recently, a genome-wide case-control study revealed that a non-synonymous single nucleotide polymorphism (SNP) (rs2230500) in a member of the protein kinase C family, PRKCH, had a significant association with lacunar infarction in 2 independent Japanese samples (OR, 1.40).32 Later, the same SNP was also found to be associated with silent lacunar infarction.33 This polymorphism increased the risk of both, ischemic and hemorrhagic stroke in the Chinese population.34 Although the study in Chinese population did not specify the subtypes of ischemic stroke, the polymorphism may be specific for cerebral SVD in Asians, because only SVD would explain increased risk of both, ischemic and hemorrhagic stroke in general. The polymorphism may also contribute to a higher proportion of both, lacunar stroke and primary ICH in the Asian population rather than Caucasians, as the minor allele is rarely found in European descendants.32
However, European large-scale collaborative studies applying GWAS technique have confirmed subtype-specific associations for cardioembolic stroke only near PITX2 and ZFHX3 and for large vessel disease at a 9p21 locus and HDAC9, but not for SVD.35 In addition, heritability estimates calculated by genome-wide complex trait analysis were 40.3% for large vessel disease, 32.6% for cardioembolic stroke but only 16.1% for SVD.36 In a meta-analysis of GWAS in Caucasian participants in 6 studies comprising the Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) consortium, the most significant association was found on chromosome 20p12 with rs2208454 for silent infarcts found on brain MRI. However, the association did not replicate in independent samples of 1,822 Caucasian and 644 non-Caucasian participants.37 As the candidate genes for cerebral SVD identified in meta-analysis have not been replicated with GWAS, they might have a limited role in lacunar stroke pathogenesis. However, the candidate gene approach can still be useful because the current GWAS cannot cover all alleles with low frequencies.
For cerebral WMH, a meta-analysis of GWAS involving 9,361 stroke-free individuals of European descent from 7 community-based cohorts identified 6 novel SNPs on chromosome 17q25, encompassing 6 known genes. Among these, the significance of two SNPs (rs3744028 and rs1055129) was replicated in an independent sample.38 However, variant alleles at these loci contributed to only a small increase in the WMH burden (4% to 8% of the overall mean WMH burden in the sample).
Recently, a GWAS with 1,545 ICH patients (664 with lobar and 881 with nonlobar) of European descent, reported a significant association between nonlobar ICH and chromosomal region 1q22 (rs2984613, OR=1.44, P=1.6×10-8), which was replicated in another dataset.39 In addition, several other SNPs within this locus also achieved genome-wide significance and some of them were listed in the GWAS report of the WMH mentioned above, which supports a common pathogenesis of both WMH and ICH.38 As described earlier, the 1425G/A SNP (rs2230500) in PRKCH was associated with an increased risk of both, ischemic stroke and hemorrhagic stroke in the Chinese population,34 but the association needs further validation in other ethnic populations.
To summarize, several important genetic loci have been identified for cerebral SVD from various ethnic backgrounds using the GWAS technique. However, detailed information on the specific genes and functional variants should be investigated further to determine the exact role of gene function or gene products in pathogenic mechanisms underlying SVD.
CADASIL is one of the most common single-gene disorders of the cerebral small blood vessels caused by mutations in the NOTCH3 gene on chromosome 19q12.40 The main clinical features include recurrent stroke, migraine, psychiatric symptoms, and progressive cognitive decline.41
Ischemic stroke is the most frequent manifestation and is present in 60% to 84% patients with CADASIL.42,43,44,45 The mean age of stroke onset is around 41 to 49 years and a majority of patients with CADASIL present with typical lacunar syndrome. Cognitive deficit is found in approximately 60% patients, with two-thirds developing dementia by age 65 years. Migraine is the most common initial symptom that occurs in 22% to 77% patients and it usually begins around age 20 years and will develop in 90% patients by age 40 years.42,43,46 Several patients have reported an aura with occasional basilar migraine, hemiplegic migraine, or migraine with prolonged aura. Psychiatric symptoms occur in 20% to 41% patients with CADASIL and mood disorders are most common.44,47 Other infrequent manifestations include epileptic seizures, acute reversible encephalopathy, and neurological complications following catheter angiography.46,48,49 Granular osmiophilic material found in the walls of affected arterioles is a pathologic hallmark of the disease, and it can be observed in skin and muscle, which is useful for pathologic diagnosis.50 Brain MRI of patients with CADASIL frequently shows progressive WMH, multiple lacunar infarcts, and microbleeding.51,52 The involvement of the anterior temporal lobe and external capsule is known to be characteristic in comparison to that in the sporadic form of WMH.53 Recently, a 7-T MRI study found a significant reduction in the number of small veins in the centrum semiovale of CADASIL patients compared with matched controls.54 This significant reduction was detected both within and outside WMH, suggesting that loss of venous vasculature may occur before the development of WMH.
The NOTCH3 gene encodes cell surface receptors that transduce signals between adjacent cells.55
NOTCH3 is mainly expressed in vascular smooth muscle cells in adults and is critical for vascular development, differentiation, and remodeling.56,57 The NOTCH3 receptor is a single-pass transmembrane protein that consists of 2,321 amino acids, and it has a large extracellular domain (ECD), with 34 tandem epidermal growth factor (EGF)-like domains. Each EGF-like domain contains 6 cysteine residues. Currently, over 170 different mutations have been reported in CADASIL and a majority of these are missense, point mutations,58 frequently located in exon 2-24, which encodes the ECD of the NOTCH3 receptor. Frequently, the mutations result in an odd number of cysteine residues added into the affected EGF. However, a small number of studies have reported that the mutations do not involve cysteine. Interestingly, common SNPs that are frequently found at the NOTCH3 gene, apart from the mutations causing CADASIL, also increase the risk of age-related WMH in individuals with hypertension.59
Although CADASIL has been reported worldwide in all ethnic groups, the exact prevalence of CADASIL is unknown. In 2002, the disease prevalence was reported to be 1.98 per 100,000 in West Scotland.60 In a study including 218 subjects with lacunar stroke, CADASIL mutation was found in only 1 patient.61 Recently, our group screened 151 consecutive patients with acute ischemic stroke for NOTCH3 mutations and found that 6 (4.0%, 95% CI, 0.9 to 7.1) had a NOTCH3 gene mutation. The prevalence of CADASIL increased to 36.0% (95% CI, 8.0 to 64.8) for patients with neuroimaging features consistent with advanced SVD.
CARASIL is a single-gene disorder of cerebral small blood vessels with autosomal recessive inheritance. This disorder is caused by mutations in the HTRA1 gene encoding HtrA serine peptidase/protease 1 (HTRA1).62,63 Since the first report of encephalopathy without hypertension from a Japanese family in 1976, the disorder has been predominantly reported in Japan.64 A small number of cases in patients from other ethnicities have also been reported recently.65,66 Although the exact prevalence is unknown, CARASIL is much rarer than CADASIL, as approximately only 50 patients have been reported so far.63
The main clinical features include early-onset lacunar stroke, progressive cognitive deficit, gait disturbance, alopecia, and low back pain.63,64 Lacunar stroke, reported in approximately 50% patients, is the most common manifestation of CARASIL and is usually found in the basal ganglia or brainstem. Cognitive deficit is the second-most common symptom and almost all patients develop dementia by 30 to 40 years of age, which is significantly earlier than CADASIL. Alopecia is the most frequent initial manifestation of the disorder, which can be found in almost 90% patients. It usually begins at adolescence and involves diffuse alopecia involving the entire scalp. Low back pain develops in approximately 80% patients and usually begins at 20 to 40 years of age. Radiologically, spondylosis deformans or disk degeneration has been identified at the cervical or thoracolumbar junction. Unlike CADASIL, migraine-like headache has not been reported with CARASIL; however, cranial MRI findings are reported to be similar to that in CADASIL patients. Pathologically, extensive degeneration of vascular smooth muscle cells and reduction in the mural extracellular matrix are found in cerebral small arteries; however, granular osmiophilic materials or amyloid deposition has not been identified.
HTRA1 is a serine protease that inhibits signaling by members of the transforming growth factor β (TGF-β) family.62 To date, 10 mutations in the HTRA1 gene have been identified in 12 families, resulting in decreased level of protease activity leading to an increase in TGF-β signaling.64 Abnormally increased TGF-β signaling seems to cause degeneration of vascular smooth muscle cells, because TGF-β plays an important role in the differentiation of vascular smooth muscle cells.
Mutations in a gene encoding type IV collagen α1 (COL4A1) were initially associated with porencephaly and infantile hemiparesis,67 however, these mutations have been later found to cause cerebral SVD even in adulthood.68,69 In a systemic review of 52 carriers of the COL4A1 mutation, stroke was present in 9 (17%) subjects in addition to infantile hemiparesis; 6 cases showed ICH and 3 cases had lacunar stroke.70 The mean age of onset of stroke was 36 years (range, 14 to 49 years), and ICH was often recurrent and provoked by minor trauma, activity, or anticoagulant use. Asymptomatic intracranial aneurysm was also found frequently, while migraine was reported in 30% cases with a mean age of onset at 30 years. In addition, several cases of systemic involvement were also found in the eye (cataracts, retinal vascular tortuosity, retinal hemorrhage, Axenfeld-Rieger anomaly), kidney (hematuria, renal cysts), and muscles (cramps). Type IV collagen is the main component of all basement membranes in humans and accordingly, mutations in COL4A1 contribute to a broad spectrum of disorders involving the brain, muscles, eyes, and kidneys. The COL4A1 gene consists of 52 exons and is located in chromosome 13q34.71 More than 30 types of mutations have been reported to date in the COL4A1 gene, and the number is rapidly increasing.72 Although COL4A1 gene mutations were known to cause instability and defects in the basement membrane, the exact mechanism linking mutations and phenotypes is presently unknown.
Hereditary vascular retinopathy, cerebroretinal vasculopathy, and hereditary endotheliopathy with retinopathy, nephropathy, and stroke (HERNS) were originally reported as distinct autosomal dominant disorders.73,74,75 In addition to vascular retinopathy as a common feature, the disorders present with various combinations of Raynaud phenomenon, migraine, cranial pseudotumor, and mild kidney or liver dysfunction. Subsequently, three disorders were mapped to a common locus in chromosome 3p21, and these are now collectively termed as autosomal dominant RVCL.76 The disorder is caused by C-terminal heterozygous, frame-shift mutations in TREX1 gene encoding a DNA-specific 3' to 5' exonuclease DNase III.77 Interestingly, homozygous mutations within this gene cause Aicardi-Goutieres syndrome, which presents as severe encephalopathy, calcification of the basal ganglia, and leukodystrophy.78 Currently, the exact mechanism underlying the mutations in TREX1 causing clinical manifestation is uncertain. The TREX1 proteins found in individuals with RVCL lack part of the C-terminus and this finding might be associated with its intracellular localization.
In a Chinese family with 11 affected patients with HERNS, progressive visual loss began at 20 to 30 years of age and other symptoms included psychiatric symptoms, migraine, and renal dysfunction.74 Neurologic deficits usually occurred as stroke-like episodes following visual loss and consisted of multifocal cortical and subcortical dysfunction. Brain MRI showed multifocal WMH lesions before neurological symptoms, which later turned into contrast-enhancing lesions with edema, frequently found at the time of focal neurologic deficit. Electron microscopic examination revealed distinctive multi-laminated vascular basement membranes in the brain and kidney, and fluorescein retinal angiography frequently showed juxtafoveolar capillary obliteration with tortuous telangiectatic microaneurysms.
FD is an X-linked, inherited disorder affecting glycosphingolipid metabolism, caused by absent or deficient lysosomal α-galactosidase A activity.79 This enzyme deficiency results in accumulation of globotriaosylceramide within the lysosomes of various organs, such as blood vessels, kidneys, heart, and dorsal root ganglia. Lysosomal α-galactosidase A is encoded by the GLA gene on chromosome Xq22, and more than 585 pathogenic mutations have been reported in the GLA gene to date. FD incidence has been estimated to be 1 per 17,000 to 117,000 in the general population,80,81 but the prevalence ranges from 0.6% to 11.1%, with an average of 4.5% in men and 3.4% in women among patients with cryptogenic stroke.82
The classic form of FD, with no detectable α-galactosidase A presents initially with angiokeratomas, acroparesthesia, hypohidrosis, corneal opacity in childhood or adolescence, and subsequently, progressive vascular disease of the heart, kidneys and brain. Burning pain that typically occurs in the extremities (acroparesthesia) is present in 60% to 80% patients, and autonomic dysfunctions such as hypohidrosis, cardiac arrhythmia, or intestinal motility disorder occurs in patients with autonomic neuropathy. Cerebrovascular complications usually develop in the fourth decade and a majority of patients with FD present with posterior circulation stroke unlike the frequency of ischemic stroke in the general population.83 In addition to SVD, patients with FD may exhibit various stroke mechanisms such as cardiogenic embolism due to ischemic heart disease or valvular heart disease, and prothrombotic state. Recurrent strokes are commonly reported, occurring in 76% to 86% patients. Elongated, ectatic, tortuous, vertebral and basilar arteries are the most common angiographic findings. Almost all patients show WMH on brain MRI at more than 55 years of age84 and the pattern of WMH does not differ significantly from that of sporadic cerebral SVD. Other evidences of cerebral SVD including lacunar infarcts, and microbleeds have also been demonstrated in patients with FD, with the so-called "pulvinar sign" on MRI, namely hyperintensity in the pulvinar on T1-weighted images, reported in 23% patients with FD.85
Since the introduction of the GWAS technique, more accurate models for risk prediction of complex disorders, based on genetic profiles are expected.86 However, genetic risk scores predicting future stroke based on genetic variants associated with stroke and its risk factors have been discouraging to date, as the models provide only a small improvement in risk prediction compared with the classical epidemiological risk factors for stroke.87,88
For rare, single-gene disorders causing cerebral SVD, timely diagnosis is important for patients and treating physicians because such patients usually have recurrent strokes that lead to worse outcomes at a young age than patients with sporadic cerebral SVD. Genetic counseling is crucial for patients and family members as part of psychological support, reproductive planning, and education of genetic testing. As effective treatments are unavailable for most of those patients at present, routine screening of asymptomatic family members is not recommended. For patients with CADASIL, conventional cerebral angiography has been reported to cause neurological complications in up to 70% patients, and therefore, other forms of angiography such as CT or magnetic resonance angiography should be considered first in patients with CADASIL.48
Because the occurrence of ICH during birth decreased dramatically with surgical delivery in an animal model for COL4A1 mutation,68 a cesarean delivery is recommended for pregnant women carrying the COL4A1 mutation to reduce birth trauma. Patients with COL4A1 mutation should also avoid other known risk factors for ICH such as head trauma or use of anticoagulants. At present, the available data suggest that enzyme replacement therapy, either agalsidase α or β, is possibly effective only for reducing neuropathic pain but not for preventing cerebrovascular complications in patients with FD.89
Although several epidemiological studies suggest the heritability of sporadic cerebral SVD, candidate gene studies have been disappointing in identifying specific genes causing the disorder. Several important genetic loci have been identified for cerebral SVD by using the GWAS technique; nevertheless, second-generation GWAS would need an even larger sample size (e.g., 100,000 to 200,000 for a complex disease such as stroke), because undiscovered susceptibility loci are expected to have smaller effect size than the loci that are known.90 To obtain such a large sample size, stroke researchers from various regions of the world have set up the International Stroke Genetics Consortium (ISGC, www.strokegenetics.org). Recently, the consortium published recommendations for phenotypic data collection, biological sample collection, and storage to maintain standardization, reliability, and quality of the data.91,92 More active participation in the international collaborative network is required from Asian countries considering the high incidence and prevalence of stroke in this region.
With the development of next-generation genotyping and sequencing platforms, DNA sequencing costs have declined dramatically in recent years,93 which would facilitate the detection of rare single-gene disorders causing cerebral SVD. In addition, the exact role of common variants within genes causing rare cerebral SVD needs further investigation, because common variants of the NOTCH3 gene show an increased risk of age-related WMH lesions among the elderly. Rare single-gene disorders can help understand the pathogenesis of the more common, sporadic form of cerebral SVD by identification of cellular, molecular, and biochemical changes underlying cerebral small vessel damage through animal models. Improved mouse models that can express the full spectrum of human cerebral SVD need to be developed to help develop targeted treatment strategies based on the pathomechanism underlying vascular damage in cerebral SVD. For example, the clearance of NOTCH3 aggregates or the reduction of mutant NOTCH3 expression could be considered as treatment for patients with CADASIL in the near future.94
References
1. Pantoni L. Cerebral small vessel disease: from pathogenesis and clinical characteristics to therapeutic challenges. Lancet Neurol 2010;9:689-701. PMID: 20610345.


2. White H, Boden-Albala B, Wang C, Elkind MS, Rundek T, Wright CB, et al. Ischemic stroke subtype incidence among whites, blacks, and Hispanics: the Northern Manhattan study. Circulation 2005;111:1327-1331. PMID: 15769776.


3. Sacco S, Marini C, Totaro R, Russo T, Cerone D, Carolei A. A population-based study of the incidence and prognosis of lacunar stroke. Neurology 2006;66:1335-1338. PMID: 16682663.


4. Chamorro A, Sacco RL, Mohr JP, Foulkes MA, Kase CS, Tatemichi TK, et al. Clinical-computed tomographic correlations of lacunar infarction in the Stroke Data Bank. Stroke 1991;22:175-181. PMID: 2003281.


5. Petty GW, Brown RD Jr, Whisnant JP, Sicks JD, O'Fallon WM, Wiebers DO. Ischemic stroke subtypes: a population-based study of incidence and risk factors. Stroke 1999;30:2513-2516. PMID: 10582970.


6. Woo D, Gebel J, Miller R, Kothari R, Brott T, Khoury J, et al. Incidence rates of first-ever ischemic stroke subtypes among blacks: a population-based study. Stroke 1999;30:2517-2522. PMID: 10582971.


7. Jung KH, Lee SH, Kim BJ, Yu KH, Hong KS, Lee BC, et al. Secular trends in ischemic stroke characteristics in a rapidly developed country: results from the Korean Stroke Registry Study (secular trends in Korean stroke). Circ Cardiovasc Qual Outcomes 2012;5:327-334. PMID: 22474244.


8. Syed NA, Khealani BA, Ali S, Hasan A, Akhtar N, Brohi H, et al. Ischemic stroke subtypes in Pakistan: the Aga Khan University Stroke Data Bank. J Pak Med Assoc 2003;53:584-588. PMID: 14765937.

9. Tsai CF, Thomas B, Sudlow CL. Epidemiology of stroke and its subtypes in Chinese vs white populations: a systematic review. Neurology 2013;81:264-272. PMID: 23858408.



10. Turin TC, Kita Y, Rumana N, Nakamura Y, Takashima N, Ichikawa M, et al. Ischemic stroke subtypes in a Japanese population: Takashima Stroke Registry, 1988-2004. Stroke 2010;41:1871-1876. PMID: 20689083.


11. Kubo M, Hata J, Doi Y, Tanizaki Y, Iida M, Kiyohara Y. Secular trends in the incidence of and risk factors for ischemic stroke and its subtypes in Japanese population. Circulation 2008;118:2672-2678. PMID: 19106389.


12. Qureshi AI, Mendelow AD, Hanley DF. Intracerebral haemorrhage. Lancet 2009;373:1632-1644. PMID: 19427958.



13. Gorelick PB, Scuteri A, Black SE, Decarli C, Greenberg SM, Iadecola C, et al. Vascular contributions to cognitive impairment and dementia: a statement for healthcare professionals from the American heart association/American stroke association. Stroke 2011;42:2672-2713. PMID: 21778438.



14. Wardlaw JM, Smith C, Dichgans M. Mechanisms of sporadic cerebral small vessel disease: insights from neuroimaging. Lancet Neurol 2013;12:483-497. PMID: 23602162.



15. Lammie GA, Brannan F, Slattery J, Warlow C. Nonhypertensive cerebral small-vessel disease. An autopsy study. Stroke 1997;28:2222-2229. PMID: 9368569.


16. Meschia JF. Ischemic stroke as a complex genetic disorder. Semin Neurol 2006;26:49-56. PMID: 16479443.



17. Flossmann E, Schulz UG, Rothwell PM. Systematic review of methods and results of studies of the genetic epidemiology of ischemic stroke. Stroke 2004;35:212-227. PMID: 14684773.


18. Polychronopoulos P, Gioldasis G, Ellul J, Metallinos IC, Lekka NP, Paschalis C, et al. Family history of stroke in stroke types and subtypes. J Neurol Sci 2002;195:117-122. PMID: 11897241.


19. Jerrard-Dunne P, Cloud G, Hassan A, Markus HS. Evaluating the genetic component of ischemic stroke subtypes: a family history study. Stroke 2003;34:1364-1369. PMID: 12714707.


20. Carmelli D, DeCarli C, Swan GE, Jack LM, Reed T, Wolf PA, et al. Evidence for genetic variance in white matter hyperintensity volume in normal elderly male twins. Stroke 1998;29:1177-1181. PMID: 9626291.


21. Atwood LD, Wolf PA, Heard-Costa NL, Massaro JM, Beiser A, D'Agostino RB, et al. Genetic variation in white matter hyperintensity volume in the Framingham Study. Stroke 2004;35:1609-1613. PMID: 15143299.


22. Woo D, Sauerbeck LR, Kissela BM, Khoury JC, Szaflarski JP, Gebel J, et al. Genetic and environmental risk factors for intracerebral hemorrhage: preliminary results of a population-based study. Stroke 2002;33:1190-1195. PMID: 11988589.


23. Tabor HK, Risch NJ, Myers RM. Candidate-gene approaches for studying complex genetic traits: practical considerations. Nat Rev Genet 2002;3:391-397. PMID: 11988764.



24. Zhang Z, Xu G, Liu D, Fan X, Zhu W, Liu X. Angiotensin-converting enzyme insertion/deletion polymorphism contributes to ischemic stroke risk: a meta-analysis of 50 case-control studies. PLoS One 2012;7:e46495PMID: 23049705.



25. Paternoster L, Chen W, Sudlow CL. Genetic determinants of white matter hyperintensities on brain scans: a systematic assessment of 19 candidate gene polymorphisms in 46 studies in 19,000 subjects. Stroke 2009;40:2020-2026. PMID: 19407234.


26. Peck G, Smeeth L, Whittaker J, Casas JP, Hingorani A, Sharma P. The genetics of primary haemorrhagic stroke, subarachnoid haemorrhage and ruptured intracranial aneurysms in adults. PLoS One 2008;3:e3691PMID: 19008959.



27. Biffi A, Sonni A, Anderson CD, Kissela B, Jagiella JM, Schmidt H, et al. Variants at APOE influence risk of deep and lobar intracerebral hemorrhage. Ann Neurol 2010;68:934-943. PMID: 21061402.



28. Biffi A, Anderson CD, Jagiella JM, Schmidt H, Kissela B, Hansen BM, et al. APOE genotype and extent of bleeding and outcome in lobar intracerebral haemorrhage: a genetic association study. Lancet Neurol 2011;10:702-709. PMID: 21741316.



29. Maxwell SS, Jackson CA, Paternoster L, Cordonnier C, Thijs V, Al-Shahi Salman R, et al. Genetic associations with brain microbleeds: Systematic review and meta-analyses. Neurology 2011;77:158-167. PMID: 21715706.



30. Yamada M, Tsukagoshi H, Otomo E, Hayakawa M. Cerebral amyloid angiopathy in the aged. J Neurol 1987;234:371-376. PMID: 3655840.



31. Pearson TA, Manolio TA. How to interpret a genome-wide association study. JAMA 2008;299:1335-1344. PMID: 18349094.


32. Kubo M, Hata J, Ninomiya T, Matsuda K, Yonemoto K, Nakano T, et al. A nonsynonymous SNP in PRKCH (protein kinase C eta) increases the risk of cerebral infarction. Nat Genet 2007;39:212-217. PMID: 17206144.


33. Serizawa M, Nabika T, Ochiai Y, Takahashi K, Yamaguchi S, Makaya M, et al. Association between PRKCH gene polymorphisms and subcortical silent brain infarction. Atherosclerosis 2008;199:340-345. PMID: 18164711.


34. Wu L, Shen Y, Liu X, Ma X, Xi B, Mi J, et al. The 1425G/A SNP in PRKCH is associated with ischemic stroke and cerebral hemorrhage in a Chinese population. Stroke 2009;40:2973-2976. PMID: 19520989.


35. Traylor M, Farrall M, Holliday EG, Sudlow C, Hopewell JC, Cheng YC, et al. Genetic risk factors for ischaemic stroke and its subtypes (the METASTROKE collaboration): a meta-analysis of genome-wide association studies. Lancet Neurol 2012;11:951-962. PMID: 23041239.



36. Bevan S, Traylor M, Adib-Samii P, Malik R, Paul NL, Jackson C, et al. Genetic heritability of ischemic stroke and the contribution of previously reported candidate gene and genomewide associations. Stroke 2012;43:3161-3167. PMID: 23042660.


37. Debette S, Bis JC, Fornage M, Schmidt H, Ikram MA, Sigurdsson S, et al. Genome-wide association studies of MRI-defined brain infarcts: meta-analysis from the CHARGE Consortium. Stroke 2010;41:210-217. PMID: 20044523.



38. Fornage M, Debette S, Bis JC, Schmidt H, Ikram MA, Dufouil C, et al. Genome-wide association studies of cerebral white matter lesion burden: the CHARGE consortium. Ann Neurol 2011;69:928-939. PMID: 21681796.



39. Woo D, Falcone GJ, Devan WJ, Brown WM, Biffi A, Howard TD, et al. Meta-analysis of genome-wide association studies identifies 1q22 as a susceptibility locus for intracerebral hemorrhage. Am J Hum Genet 2014;94:511-521. PMID: 24656865.



40. Joutel A, Corpechot C, Ducros A, Vahedi K, Chabriat H, Mouton P, et al. Notch3 mutations in CADASIL, a hereditary adult-onset condition causing stroke and dementia. Nature 1996;383:707-710. PMID: 8878478.



41. Choi JC. Cerebral autosomal dominant arteriopathy with subcortical infarcts and leukoencephalopathy: a genetic cause of cerebral small vessel disease. J Clin Neurol 2010;6:1-9. PMID: 20386637.



42. Chabriat H, Vahedi K, Iba-Zizen MT, Joutel A, Nibbio A, Nagy TG, et al. Clinical spectrum of CADASIL: a study of 7 families. Cerebral autosomal dominant arteriopathy with subcortical infarcts and leukoencephalopathy. Lancet 1995;346:934-939. PMID: 7564728.


43. Dichgans M, Mayer M, Uttner I, Bruning R, Muller-Hocker J, Rungger G, et al. The phenotypic spectrum of CADASIL: clinical findings in 102 cases. Ann Neurol 1998;44:731-739. PMID: 9818928.


44. Desmond DW, Moroney JT, Lynch T, Chan S, Chin SS, Mohr JP. The natural history of CADASIL: a pooled analysis of previously published cases. Stroke 1999;30:1230-1233. PMID: 10356105.


45. Choi JC, Song SK, Lee JS, Kang SY, Kang JH. Diversity of stroke presentation in CADASIL: study from patients harboring the predominant NOTCH3 mutation R544C. J Stroke Cerebrovasc Dis 2013;22:126-131. PMID: 21852154.


46. Singhal S, Bevan S, Barrick T, Rich P, Markus HS. The influence of genetic and cardiovascular risk factors on the CADASIL phenotype. Brain 2004;127:2031-2038. PMID: 15229130.



47. Valenti R, Poggesi A, Pescini F, Inzitari D, Pantoni L. Psychiatric disturbances in CADASIL: a brief review. Acta Neurol Scand 2008;118:291-295. PMID: 18384453.


48. Dichgans M, Petersen D. Angiographic complications in CADASIL. Lancet 1997;349:776-777. PMID: 9074584.

49. Feuerhake F, Volk B, Ostertag CB, Jungling FD, Kassubek J, Orszagh M, et al. Reversible coma with raised intracranial pressure: an unusual clinical manifestation of CADASIL. Acta Neuropathol 2002;103:188-192. PMID: 11810186.


50. Chabriat H, Joutel A, Dichgans M, Tournier-Lasserve E, Bousser MG. Cadasil. Lancet Neurol 2009;8:643-653. PMID: 19539236.


51. van den Boom R, Lesnik Oberstein SA, Ferrari MD, Haan J, van Buchem MA. Cerebral autosomal dominant arteriopathy with subcortical infarcts and leukoencephalopathy: MR imaging findings at different ages--3rd-6th decades. Radiology 2003;229:683-690. PMID: 14593195.


52. Chabriat H, Levy C, Taillia H, Iba-Zizen MT, Vahedi K, Joutel A, et al. Patterns of MRI lesions in CADASIL. Neurology 1998;51:452-457. PMID: 9710018.


53. O'Sullivan M, Jarosz JM, Martin RJ, Deasy N, Powell JF, Markus HS. MRI hyperintensities of the temporal lobe and external capsule in patients with CADASIL. Neurology 2001;56:628-634. PMID: 11245715.


54. De Guio F, Vignaud A, Ropele S, Duering M, Duchesnay E, Chabriat H, et al. Loss of venous integrity in cerebral small vessel disease: a 7-T MRI study in cerebral autosomal-dominant arteriopathy with subcortical infarcts and leukoencephalopathy (CADASIL). Stroke 2014;45:2124-2126. PMID: 24867926.


55. Artavanis-Tsakonas S, Rand MD, Lake RJ. Notch signaling: cell fate control and signal integration in development. Science 1999;284:770-776. PMID: 10221902.


56. Joutel A, Andreux F, Gaulis S, Domenga V, Cecillon M, Battail N, et al. The ectodomain of the Notch3 receptor accumulates within the cerebrovasculature of CADASIL patients. J Clin Invest 2000;105:597-605. PMID: 10712431.



57. Alva JA, Iruela-Arispe ML. Notch signaling in vascular morphogenesis. Curr Opin Hematol 2004;11:278-283. PMID: 15314528.


58. Tikka S, Mykkanen K, Ruchoux MM, Bergholm R, Junna M, Poyhonen M, et al. Congruence between NOTCH3 mutations and GOM in 131 CADASIL patients. Brain 2009;132:933-939. PMID: 19174371.




59. Schmidt H, Zeginigg M, Wiltgen M, Freudenberger P, Petrovic K, Cavalieri M, et al. Genetic variants of the NOTCH3 gene in the elderly and magnetic resonance imaging correlates of age-related cerebral small vessel disease. Brain 2011;134:3384-3397. PMID: 22006983.




60. Razvi SS, Davidson R, Bone I, Muir KW. The prevalence of cerebral autosomal dominant arteriopathy with subcortical infarcts and leucoencephalopathy (CADASIL) in the west of Scotland. J Neurol Neurosurg Psychiatry 2005;76:739-741. PMID: 15834040.



61. Dong Y, Hassan A, Zhang Z, Huber D, Dalageorgou C, Markus HS. Yield of screening for CADASIL mutations in lacunar stroke and leukoaraiosis. Stroke 2003;34:203-205. PMID: 12511775.


62. Hara K, Shiga A, Fukutake T, Nozaki H, Miyashita A, Yokoseki A, et al. Association of HTRA1 mutations and familial ischemic cerebral small-vessel disease. N Engl J Med 2009;360:1729-1739. PMID: 19387015.


63. Fukutake T. Cerebral autosomal recessive arteriopathy with subcortical infarcts and leukoencephalopathy (CARASIL): from discovery to gene identification. J Stroke Cerebrovasc Dis 2011;20:85-93. PMID: 21215656.


64. Nozaki H, Nishizawa M, Onodera O. Features of cerebral autosomal recessive arteriopathy with subcortical infarcts and leukoencephalopathy. Stroke 2014;45:3447-3457. PMID: 25116877.


65. Bianchi S, Di Palma C, Gallus GN, Taglia I, Poggiani A, Rosini F, et al. Two novel HTRA1 mutations in a European CARASIL patient. Neurology 2014;82:898-900. PMID: 24500651.


66. Mendioroz M, Fernandez-Cadenas I, Del Rio-Espinola A, Rovira A, Sole E, Fernandez-Figueras MT, et al. A missense HTRA1 mutation expands CARASIL syndrome to the Caucasian population. Neurology 2010;75:2033-2035. PMID: 21115960.


67. Gould DB, Phalan FC, Breedveld GJ, van Mil SE, Smith RS, Schimenti JC, et al. Mutations in Col4a1 cause perinatal cerebral hemorrhage and porencephaly. Science 2005;308:1167-1171. PMID: 15905400.


68. Gould DB, Phalan FC, van Mil SE, Sundberg JP, Vahedi K, Massin P, et al. Role of COL4A1 in small-vessel disease and hemorrhagic stroke. N Engl J Med 2006;354:1489-1496. PMID: 16598045.


69. Sibon I, Coupry I, Menegon P, Bouchet JP, Gorry P, Burgelin I, et al. COL4A1 mutation in Axenfeld-Rieger anomaly with leukoencephalopathy and stroke. Ann Neurol 2007;62:177-184. PMID: 17696175.


70. Lanfranconi S, Markus HS. COL4A1 mutations as a monogenic cause of cerebral small vessel disease: a systematic review. Stroke 2010;41:e513-e518. PMID: 20558831.


71. Emanuel BS, Sellinger BT, Gudas LJ, Myers JC. Localization of the human procollagen alpha 1(IV) gene to chromosome 13q34 by in situ hybridization. Am J Hum Genet 1986;38:38-44. PMID: 3753820.


72. Kuo DS, Labelle-Dumais C, Gould DB. COL4A1 and COL4A2 mutations and disease: insights into pathogenic mechanisms and potential therapeutic targets. Hum Mol Genet 2012;21:R97-R110. PMID: 22914737.




73. Grand MG, Kaine J, Fulling K, Atkinson J, Dowton SB, Farber M, et al. Cerebroretinal vasculopathy. A new hereditary syndrome. Ophthalmology 1988;95:649-659. PMID: 3174024.


74. Jen J, Cohen AH, Yue Q, Stout JT, Vinters HV, Nelson S, et al. Hereditary endotheliopathy with retinopathy, nephropathy, and stroke (HERNS). Neurology 1997;49:1322-1330. PMID: 9371916.


75. Terwindt GM, Haan J, Ophoff RA, Groenen SM, Storimans CW, Lanser JB, et al. Clinical and genetic analysis of a large Dutch family with autosomal dominant vascular retinopathy, migraine and Raynaud's phenomenon. Brain 1998;121(Pt 2):303-316. PMID: 9549508.



76. Ophoff RA, DeYoung J, Service SK, Joosse M, Caffo NA, Sandkuijl LA, et al. Hereditary vascular retinopathy, cerebroretinal vasculopathy, and hereditary endotheliopathy with retinopathy, nephropathy, and stroke map to a single locus on chromosome 3p21.1-p21.3. Am J Hum Genet 2001;69:447-453. PMID: 11438888.



77. Richards A, van den Maagdenberg AM, Jen JC, Kavanagh D, Bertram P, Spitzer D, et al. C-terminal truncations in human 3'-5' DNA exonuclease TREX1 cause autosomal dominant retinal vasculopathy with cerebral leukodystrophy. Nat Genet 2007;39:1068-1070. PMID: 17660820.


78. Crow YJ, Hayward BE, Parmar R, Robins P, Leitch A, Ali M, et al. Mutations in the gene encoding the 3'-5' DNA exonuclease TREX1 cause Aicardi-Goutieres syndrome at the AGS1 locus. Nat Genet 2006;38:917-920. PMID: 16845398.



79. Brady RO. Enzymatic abnormalities in diseases of sphingolipid metabolism. Clin Chem 1967;13:565-577. PMID: 5006481.


80. Houge G, Skarbovik AJ. Fabry disease--a diagnostic and therapeutic challenge. Tidsskr Nor Laegeforen 2005;125:1004-1006. PMID: 15852071.

81. Meikle PJ, Hopwood JJ, Clague AE, Carey WF. Prevalence of lysosomal storage disorders. JAMA 1999;281:249-254. PMID: 9918480.


82. Shi Q, Chen J, Pongmoragot J, Lanthier S, Saposnik G. Prevalence of Fabry disease in stroke patients--a systematic review and meta-analysis. J Stroke Cerebrovasc Dis 2014;23:985-992. PMID: 24126289.


83. Mitsias P, Levine SR. Cerebrovascular complications of Fabry's disease. Ann Neurol 1996;40:8-17. PMID: 8687196.


84. Crutchfield KE, Patronas NJ, Dambrosia JM, Frei KP, Banerjee TK, Barton NW, et al. Quantitative analysis of cerebral vasculopathy in patients with Fabry disease. Neurology 1998;50:1746-1749. PMID: 9633721.


85. Moore DF, Ye F, Schiffmann R, Butman JA. Increased signal intensity in the pulvinar on T1-weighted images: a pathognomonic MR imaging sign of Fabry disease. AJNR Am J Neuroradiol 2003;24:1096-1101. PMID: 12812932.


86. Kraft P, Wacholder S, Cornelis MC, Hu FB, Hayes RB, Thomas G, et al. Beyond odds ratios--communicating disease risk based on genetic profiles. Nat Rev Genet 2009;10:264-269. PMID: 19238176.



87. Malik R, Bevan S, Nalls MA, Holliday EG, Devan WJ, Cheng YC, et al. Multilocus genetic risk score associates with ischemic stroke in case-control and prospective cohort studies. Stroke 2014;45:394-402. PMID: 24436234.



88. Ibrahim-Verbaas CA, Fornage M, Bis JC, Choi SH, Psaty BM, Meigs JB, et al. Predicting stroke through genetic risk functions: the CHARGE Risk Score Project. Stroke 2014;45:403-412. PMID: 24436238.



89. El Dib RP, Nascimento P, Pastores GM. Enzyme replacement therapy for Anderson-Fabry disease. Cochrane Database Syst Rev 2013;2:CD006663PMID: 23450571.
90. Park JH, Wacholder S, Gail MH, Peters U, Jacobs KB, Chanock SJ, et al. Estimation of effect size distribution from genome-wide association studies and implications for future discoveries. Nat Genet 2010;42:570-575. PMID: 20562874.




91. Battey TW, Valant V, Kassis SB, Kourkoulis C, Lee C, Anderson CD, et al. Recommendations From the International Stroke Genetics Consortium, Part 2: Biological Sample Collection and Storage. Stroke 2015;46:285-290. PMID: 25492904.



92. Majersik JJ, Cole JW, Golledge J, Rost NS, Chan YF, Gurol ME, et al. Recommendations From the International Stroke Genetics Consortium, Part 1: Standardized Phenotypic Data Collection. Stroke 2015;46:279-284. PMID: 25492903.



93. Wetterstrand K. DNA Sequencing Costs: Data from the NHGRI Genome Sequencing Program (GSP) Accessed. December 11, 2014. Available at: www.genome.gov/sequencingcosts.
94. Joutel A, Faraci FM. Cerebral small vessel disease: insights and opportunities from mouse models of collagen IV-related small vessel disease and cerebral autosomal dominant arteriopathy with subcortical infarcts and leukoencephalopathy. Stroke 2014;45:1215-1221. PMID: 24503668.



Table 2
Characteristics of single-gene disorders causing cerebral SVD
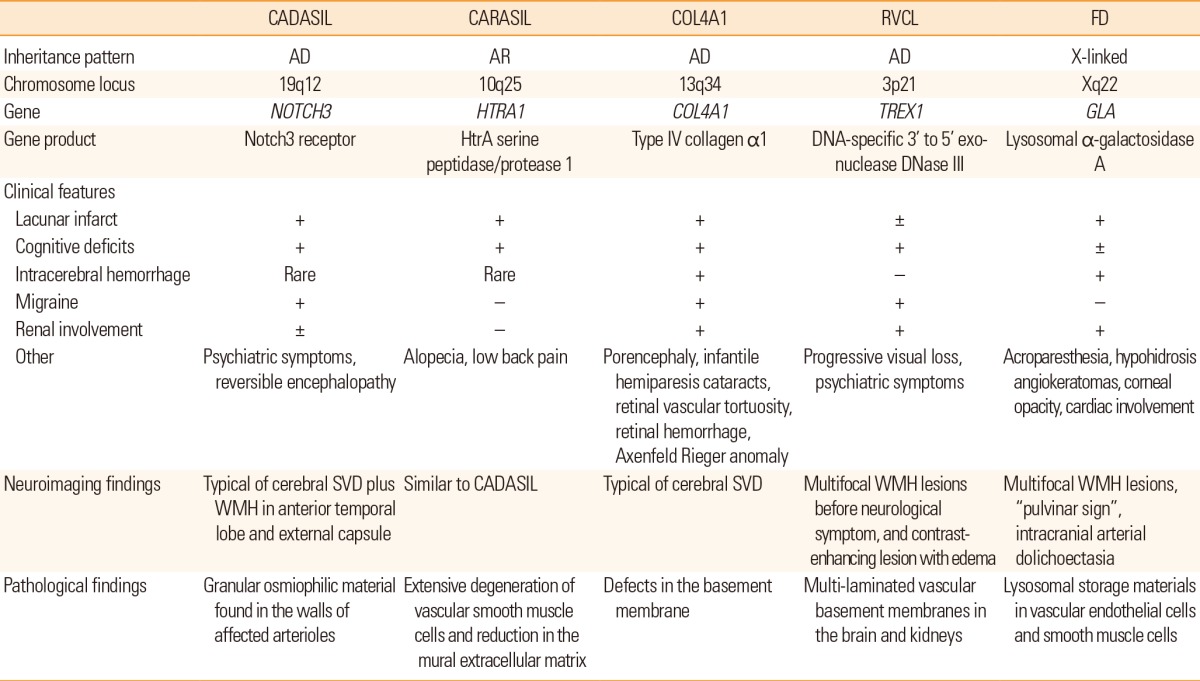
CADASIL, Cerebral autosomal dominant arteriopathy with subcortical infarcts and leukoencephalopathy; CARASIL, Cerebral autosomal recessive arteriopathy with subcortical infarcts and leukoencephalopathy; RVCL, retinal vasculopathy with cerebral leukodystrophy; FD, Fabry disease; AD, autosomal dominant; AR, autosomal recessive; SVD, small vessel disease; WMH, white matter hyperintensities.